The Beddington Medal is the BSDB’s major commendation to promising young biologists, awarded for the best PhD thesis in Developmental Biology defended in the year previous to the award. Rosa Beddington was one of the greatest talents and inspirational leaders in the field of developmental biology. Rosa made an enormous contribution to the field in general and to the BSDB in particular, so it seemed entirely appropriate that the Society should establish a lasting memorial to her. The design of the medal, mice on a stylised DNA helix, is from artwork by Rosa herself.
Like many years, it was a tough decision for the BSBD committee to choose a winner for the 2025 Beddington medal. We are pleased to announce that this goes to Rory Maizels, for his PhD work at the Crick Institute on differential signal interpretation and cell fate decisions in the developing neural tube.

 I am writing to enthusiastically support Rory Maizels’s nomination for the Beddington Medal. His PhD work represents a remarkable achievement that advances our field’s long-standing goal: developing dynamical models that capture the full complexity of developmental systems. The central challenge in developmental biology is to understand how complex, multicellular tissues emerge from the coordinated actions of individual cells. While we have made great strides in identifying key molecular players and mapping gene regulatory networks, we still lack the ability to create predictive dynamical models that capture development in its full complexity. Rory’s work represents a critical step toward addressing this fundamental challenge. What sets Rory’s contribution apart is both its comprehensive scope and meticulous execution. Rather than pursuing flashy but superficial advances, he focused on building robust foundations – developing and rigorously validating new experimental and computational approaches that together enable dynamic modelling of development at scale. Remarkably, Rory personally drove every aspect of the project: from optimising molecular biology protocols and establishing automated laboratory workflows, to designing novel machine learning frameworks for analysing the resulting data. This rare combination of experimental and computational expertise allowed him to iterate between theory and practice in a uniquely effective way.
I am writing to enthusiastically support Rory Maizels’s nomination for the Beddington Medal. His PhD work represents a remarkable achievement that advances our field’s long-standing goal: developing dynamical models that capture the full complexity of developmental systems. The central challenge in developmental biology is to understand how complex, multicellular tissues emerge from the coordinated actions of individual cells. While we have made great strides in identifying key molecular players and mapping gene regulatory networks, we still lack the ability to create predictive dynamical models that capture development in its full complexity. Rory’s work represents a critical step toward addressing this fundamental challenge. What sets Rory’s contribution apart is both its comprehensive scope and meticulous execution. Rather than pursuing flashy but superficial advances, he focused on building robust foundations – developing and rigorously validating new experimental and computational approaches that together enable dynamic modelling of development at scale. Remarkably, Rory personally drove every aspect of the project: from optimising molecular biology protocols and establishing automated laboratory workflows, to designing novel machine learning frameworks for analysing the resulting data. This rare combination of experimental and computational expertise allowed him to iterate between theory and practice in a uniquely effective way.
Prior to his PhD, Rory built a strong foundation through diverse research experience: molecular biology at LMCB UCL, developing computational tools for mitochondrial research at Oxford (resulting in an eLife publication), and completing the prestigious Frank Knox Fellowship at Harvard in Computational Science and Engineering. It is important to emphasise that this fellowship was not simply a bioinformatics MSc but a computational course aimed at engineers and data scientists. This unique background prepared him perfectly for tackling the emerging challenges in single-cell genomics and developmental biology. At the Crick, he quickly demonstrated exceptional independence and scientific maturity, showing deep knowledge of the field while working autonomously and effectively communicating complex ideas to others.
In the first months in the lab (during the COVID pandemic), Rory led the computational analysis of a major single-cell RNA sequencing study of human neural development, analysing data from multiple stages of embryonic spinal cord tissue to identify distinct cell types and map differentiation pathways. His analysis not only revealed the diversity of neural cell types and their developmental trajectories but also provided important comparative insights between human and mouse development, demonstrating both his technical capabilities and his ability to collaborate effectively on complex projects. This work is published.
In his main PhD project, Rory developed novel experimental and computational methods. This delivered three major technical innovations that together advance our ability to study developmental dynamics. First, he developed sci-FATE2, an optimized and semi-automated protocol for metabolic labelling and single-cell RNA sequencing that matches commercial platforms in quality while being simpler to implement. This is published as a methods paper. Second, he created Velvet, a deep learning framework that improves upon existing methods for inferring cell state transitions from RNA data by integrating neighbourhood information into its velocity calculations. Finally, he extended this work with VelvetSDE, a cutting-edge neural stochastic differential equation system that can predict long-term cell fate trajectories and identify key decision points in development, while capturing the inherent variability in cellular decision-making. Applying this to data from the neural tube led to the realisation that expression of Shh modulators are crucial for differential signal interpretation and cell fate decision in the developing neural tube. This combination of experimental and computational advances provides a robust framework for studying the complex dynamics of development at unprecedented scale and resolution. The work recasts single-cell analyses from descriptions of observed data to models of the dynamics that generated them, providing a framework for investigating developmental fate decisions. This work is published.
Rory’s unique combination of creativity, determination, and technical expertise is responsible for the success of the project. His exceptional strengths in both experimental and computational approaches, spanning molecular biology to machine learning, gives him an ability to tackle complex biological problems from multiple angles. But his ability is not limited to technical skills. He is a deep thinker and has developed a clear and far-reaching view of the future of developmental biology. These scholarly capabilities are evidenced by his invited review on single-cell transcriptomics, which he authored independently following a well-received presentation at the Royal Society. We are also completing an article that sets out a vision for developmental biology in the single cell genomics era. In short, Rory is both a thinker and a doer.
The impact of Rory’s work is already evident in the catalytic effect it is having in the field. It has attracted substantial funding (three grants: CRUK Development, Crick I2I Funding, BBSRC project grant) and underpins five new projects in the lab, including single-cell screening of glioma transcription factors, timeresolved sequencing of organoids, and targeted sequencing approaches. Beyond our group, it has enabled new collaborations in cancer screening, neurodegeneration research, and immunology with leading labs. Most notably, this work formed the foundation for Rory’s successful fellowship application for post-doc at EBI and Sanger, where he will further develop these approaches.
Rory exemplifies the qualities we hope to cultivate in our field: deep theoretical understanding combined with practical capability, rigorous methodology alongside creative vision, and the ability to both conceive and execute transformative research. He is not just technically accomplished but a profound thinker about the future of developmental biology and a clear communicator. His work demonstrates both the insight to identify fundamental challenges and the skill to address them systematically.
Given the extraordinary breadth and depth of his contributions, his proven ability to execute complex interdisciplinary projects, and the clear impact his work is already having on the field, I believe Rory Maizels is an outstanding recipient of the Beddington Medal. He represents the kind of scientist who will help lead our field into its next phase, where we can finally begin to build a comprehensive understanding of development.
James Briscoe
Papers:
Maizels, R. J., and Briscoe, J. (2025). Gene regulatory networks: from correlaCve models to causal explanaCons. In prepara(on.
Maizels, R. J. (2024). A dynamical perspecCve: moving towards mechanism in single-cell transcriptomics. Philos. Trans. R. Soc. B
Maizels, R. J., Snell, D. M., and Briscoe, J. (2024). ReconstrucCng developmental trajectories using latent dynamical systems and Cme-resolved transcriptomics. Cell Systems
Maizels, R. J., Snell, D. M., and Briscoe, J. (2024). A protocol for Cme-resolved transcriptomics through metabolic labeling and combinatorial indexing. STAR Protocols
Rayon, T., Maizels, R. J., Barrington, C., and Briscoe, J. (2021). Single-cell transcriptome profiling of the human developing spinal cord reveals a conserved genetic programme with human-specific features. Development

 We are very happy to announce that this year’s winner of the BSDB Wolpert medal is Prof. Sally Lowell from the University of Edinburgh.
We are very happy to announce that this year’s winner of the BSDB Wolpert medal is Prof. Sally Lowell from the University of Edinburgh. Until a few months ago, Sally was our BSDB meeting secretary and she was outstanding in this role, bringing many new initiatives including the BSDB childcare and disability travel awards. She pushed hard in many ways for diversity, inclusivity and sustainability, so that during her tenure the BSDB became one of the leading drivers of new ways of running conferences. This climaxed with our recent hosting of the European Dev Biol Congress where she pushed for – and made work – a programme made up largely of ECR speakers from across Europe, with a unique three hub arrangement with interdigitating talks beamed in from Paris and Barcelona to the central host hub of Oxford. This was a pioneering “experiment” that could have gone badly wrong, but instead worked exceptionally well, and will have set a precedent for others to follow. Several colleagues from sister dev biol societies across Europe congratulated us on how brave the BSDB was to run such a meeting and how successful it had been. This kudos for the BSDB was largely down to Sally.
Until a few months ago, Sally was our BSDB meeting secretary and she was outstanding in this role, bringing many new initiatives including the BSDB childcare and disability travel awards. She pushed hard in many ways for diversity, inclusivity and sustainability, so that during her tenure the BSDB became one of the leading drivers of new ways of running conferences. This climaxed with our recent hosting of the European Dev Biol Congress where she pushed for – and made work – a programme made up largely of ECR speakers from across Europe, with a unique three hub arrangement with interdigitating talks beamed in from Paris and Barcelona to the central host hub of Oxford. This was a pioneering “experiment” that could have gone badly wrong, but instead worked exceptionally well, and will have set a precedent for others to follow. Several colleagues from sister dev biol societies across Europe congratulated us on how brave the BSDB was to run such a meeting and how successful it had been. This kudos for the BSDB was largely down to Sally. The
The  Originally trained as an engineer and physicist, JP Vincent became a developmental biologist by accident, when his PhD advisor George Oster, a mechanical engineer turned biologist, suggested that he look at the fluid dynamics of Xenopus eggs. He was lucky to be hosted by John Gerhart for the wet part of this project and was quickly taken by the warmth of the developmental biology community and the range of questions that developmental biology addresses. Since then, JP has been inspired by classical questions of developmental biology such as axis formation, cell fate determination, morphogen gradient formation and tissue renewal, and strived to bring methods from other disciplines to address them. His work has questioned established dogma, uncovered new mechanisms, and brought outsiders into the developmental biology field.
Originally trained as an engineer and physicist, JP Vincent became a developmental biologist by accident, when his PhD advisor George Oster, a mechanical engineer turned biologist, suggested that he look at the fluid dynamics of Xenopus eggs. He was lucky to be hosted by John Gerhart for the wet part of this project and was quickly taken by the warmth of the developmental biology community and the range of questions that developmental biology addresses. Since then, JP has been inspired by classical questions of developmental biology such as axis formation, cell fate determination, morphogen gradient formation and tissue renewal, and strived to bring methods from other disciplines to address them. His work has questioned established dogma, uncovered new mechanisms, and brought outsiders into the developmental biology field.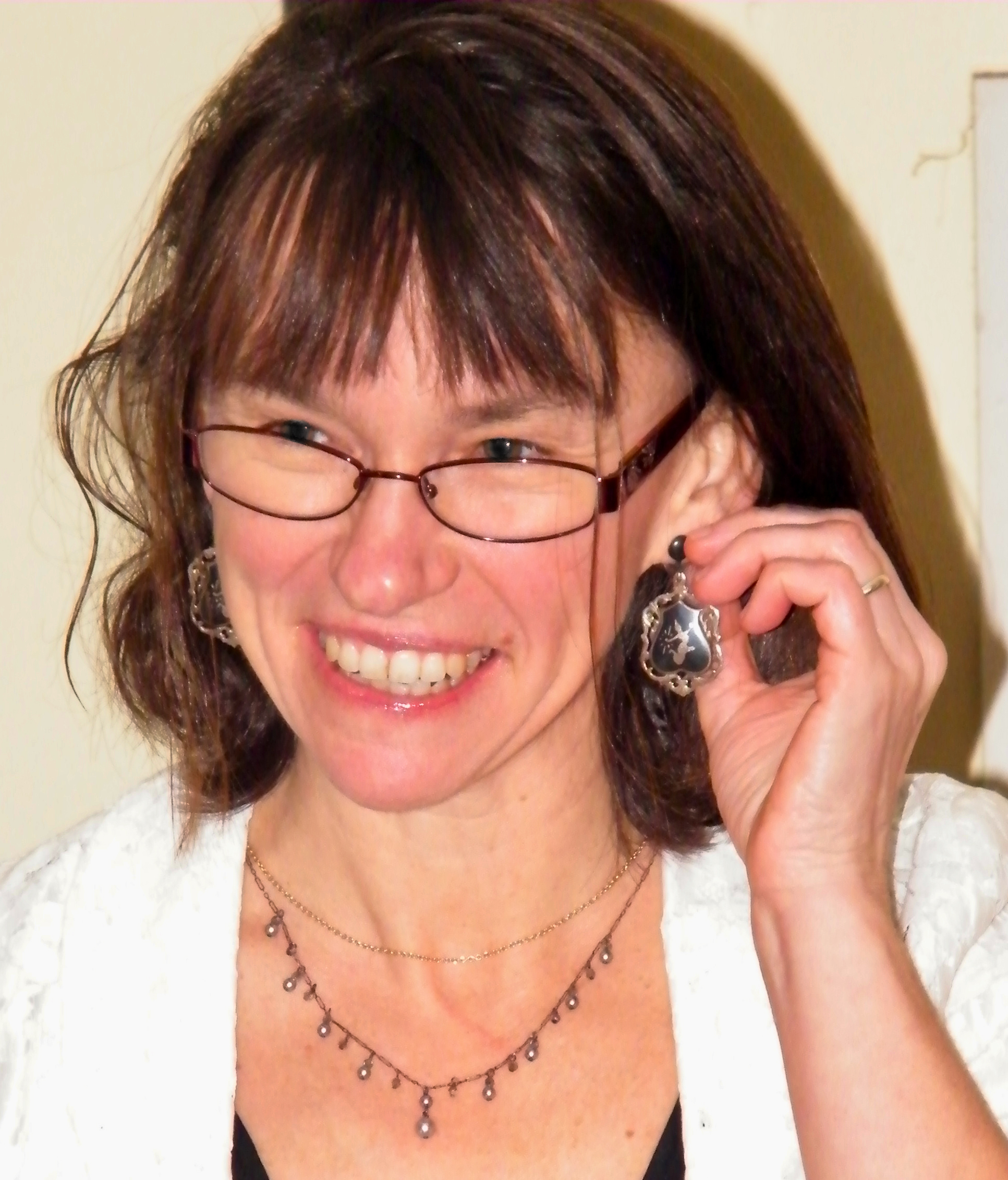
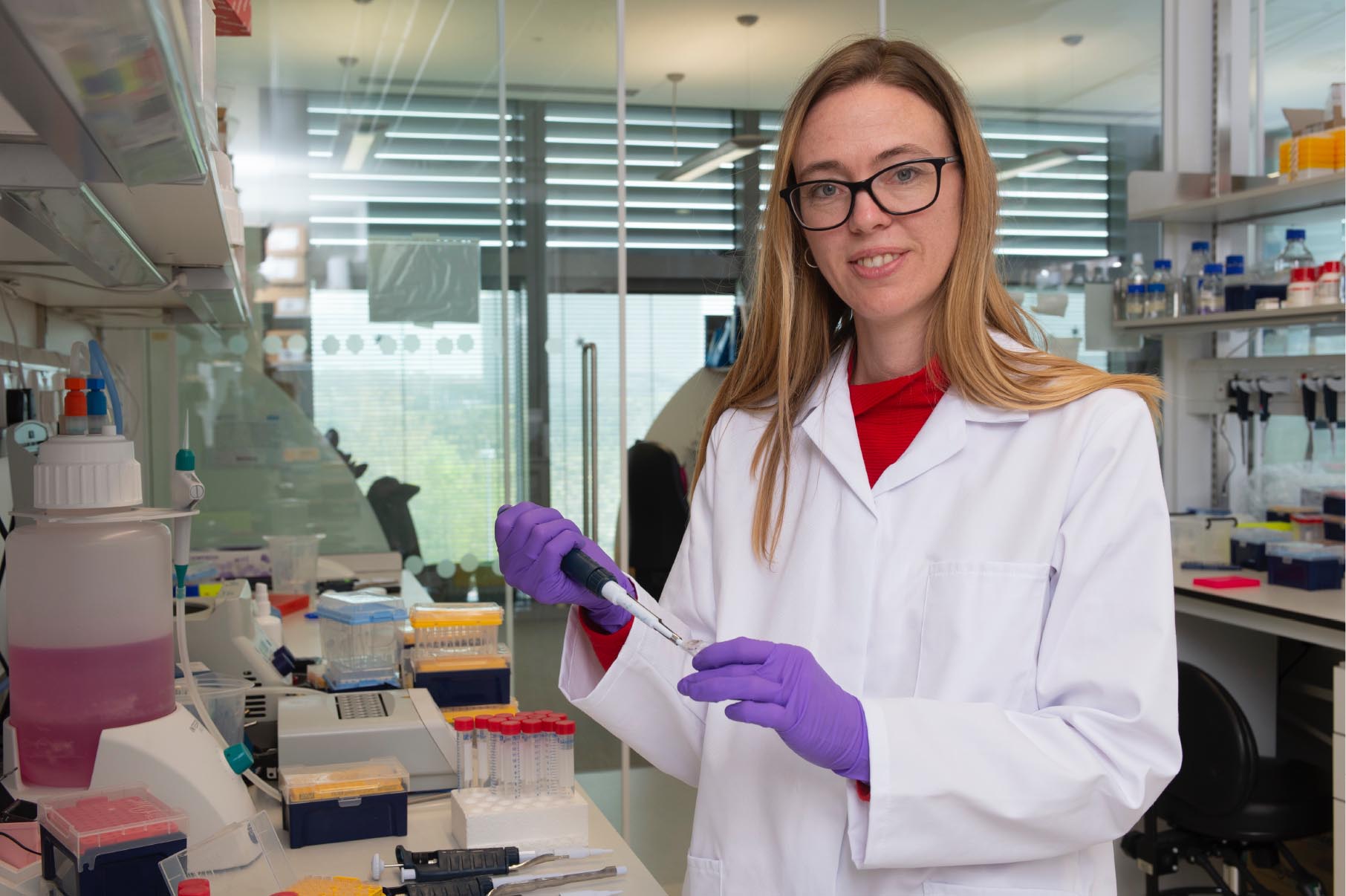
 Madeline is a most worthy recipient of the Tickle Medal. Madeline started her independent laboratory in 2015 at the Medical Research Council Laboratory of Molecular Biology (University of Cambridge) following landmark and courageous work developing organoids for the most complex and inaccessible of organs – the human brain. To do so, Madeline, looked to Developmental Biology to rationally decide upon conditions that might guide cellular self-organisation into variations of this organ, or regions or aspects of it (e.g., mimicking different axial levels and stages, more recently capable of secreting cerebrospinal fluid). Although there is not an embryo in sight, Madeline’s work has provided unprecedented functional access to models and perturbations relevant to understanding human brain development. It has also allowed probing of likely mechanisms of brain evolution and indeed its marriage with development, in the field of “evo-devo”. Finally, it has allowed investigation of intersections between development and human disease. Under her guidance, iterations of healthy and disease modelling brain organoids are contributing a wealth of what we could call equally “cellular synthetic biology” or “engineered developmental biology”. We are learning what it takes – at the molecular, cell biological, and supra-cellular levels – to coax cells into building particular fate and morphological ensembles that recapitulate important aspects of brain development.
Madeline is a most worthy recipient of the Tickle Medal. Madeline started her independent laboratory in 2015 at the Medical Research Council Laboratory of Molecular Biology (University of Cambridge) following landmark and courageous work developing organoids for the most complex and inaccessible of organs – the human brain. To do so, Madeline, looked to Developmental Biology to rationally decide upon conditions that might guide cellular self-organisation into variations of this organ, or regions or aspects of it (e.g., mimicking different axial levels and stages, more recently capable of secreting cerebrospinal fluid). Although there is not an embryo in sight, Madeline’s work has provided unprecedented functional access to models and perturbations relevant to understanding human brain development. It has also allowed probing of likely mechanisms of brain evolution and indeed its marriage with development, in the field of “evo-devo”. Finally, it has allowed investigation of intersections between development and human disease. Under her guidance, iterations of healthy and disease modelling brain organoids are contributing a wealth of what we could call equally “cellular synthetic biology” or “engineered developmental biology”. We are learning what it takes – at the molecular, cell biological, and supra-cellular levels – to coax cells into building particular fate and morphological ensembles that recapitulate important aspects of brain development.